# upload recreational landings
rec <- read.csv("data/recLandings.csv")
names(rec)[1] <- "Year"
# summarize by basin and total
rec$ATL <- rowSums(rec[3:6], na.rm = T)
rec$ALL <- rec$GULF + rec$ATL
# consider 1990 and forward
rec <- rec[which(rec$Year >= 1990), ]
# sort years from lowest to higher landings
sortyr <- rec$Year[order(rec$ALL)]
# cols <- rainbow(100, start = 0.1, end = 0.65)[100:1]
cols <- c(0, rainbow(3, start = 0.4, end = 0.6), 2)
# set up plot
nf <- layout(matrix(c(2:56, rep(1, 11), 57:111), 11, 11, byrow = FALSE),
c(rep(3, 5), 2, rep(3, 5)), c(4, rep(3, 11)))
#layout.show(nf)
#legend
par(mar = c(0, 0, 0, 0))
plot(1, type = "n", axes = FALSE, xlab = "", ylab = "")
legend("center", fill = cols, c("<26", "26-27", "27-28.5", "28.5-30", ">30"),
border = c(1, 0, 0, 0, 0),
pt.cex = 2, bty = "n",
title = "SST\n(deg C)", title.font = 2)
# sort years from lowest to highest landings
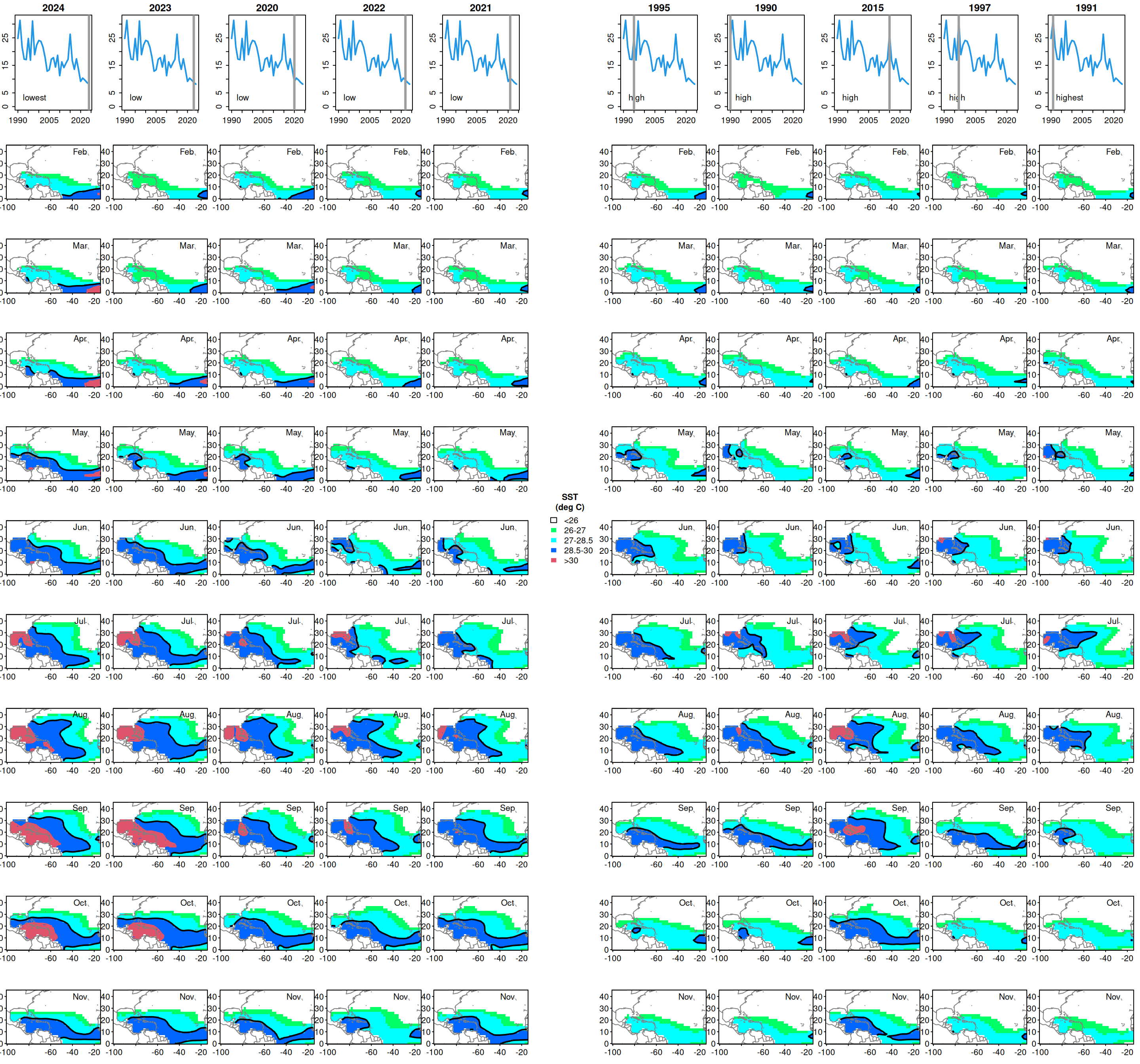
yrs <- c(sortyr[1:5], sortyr[(length(sortyr)-4): length(sortyr)])
#yrs
status <- c(rep("low", 5), rep("high", 5))
status[1] <- "lowest"
status[10] <- "highest"
# for lowest and highest years, visualize temperature patterns across the Atlantic
for (j in 1: length(yrs)) {
k <- which(yr == yrs[j])
# print(data.frame(tim[k], yr[k], mon[k]))
par(mex = 0.5, mar = c(2, 2, 2, 2)+1)
plot(rec$Year, rec$ALL/10^6, type = "l", col = 4, ylim = c(0, 32), lwd = 2, ylab = "rec landings",
main = yrs[j])
text(1990, 3, status[j], pos = 4)
abline(v = yrs[j], col = 8, lwd = 3)
# plot months February through November
for (i in k[2:11]) {
par(mex = 0.3, mar = c(2, 3, 2, 0))
map("world", xlim = c(-100, -15), ylim = c(0, 45), col = 0)
axis(1); axis(2, las = 2); box()
image(lon, lat, sst[, , i], add = T, col = cols, breaks = c(-2, 26, 27, 28.5, 30, 32))
text(-33, 40, paste(month.abb[mon[i]]), cex = 1)
map("world", add = T, col = gray(0.5)); box()
contour(lon, lat, sst[, , i], levels = c(30), add = T, col = c(2), lwd = 2, drawlabels = FALSE)
contour(lon, lat, sst[, , i], levels = c(28.5), add = T, col = 1, lwd = 2, drawlabels = FALSE)
}
}